Evolution
Vertebrate biology is distinct from that of invertebrates, plants and prokaryotes. It therefore follows that the means in which we respond to viruses is also distinct. In the tenOever lab, we are interested in learning how cellular antiviral defenses are shaped over time and exploiting these mechanisms to generate new technologies in an effort to create novel therapeutics.
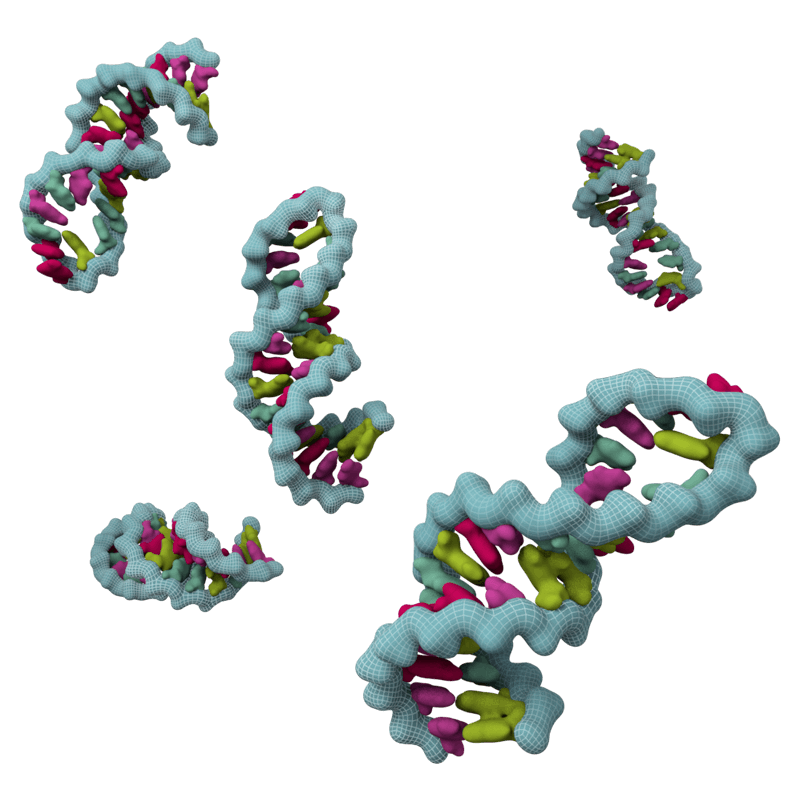
The emergence of microRNA (miRNA) biology coincided with multicellularity and arose independently in plants and animals. Similar to the antiviral defenses of plants, worms, and arthropods, most eukaryotic species utilize miRNAs as a means of imposing post transcriptional silencing. This type of regulation is distinct from the antiviral RNAi pathways observed in plants, worms, and arthropods. Unlike the biology used in antiviral defenses, these small RNAs are genome-encoded and interact with mRNA with imperfect complementarity. This difference prevents small RNA-mediated cleavage by the RNA induced silencing complex and, as a result, these miRNAs are considered fine-tuners of host biology. However, should one introduce a perfect binding site for a given miRNA into a desired transcript, cleavage and potent silencing can be restored.
We first adapted this idea as a vaccine platform where we could design a virus to be attenuated in one species but replicate to high titers in another based on the exploitation of host-specific miRNA profiles. Building on this idea, we later adapted it as a means of molecular biocontainment to generate viruses that could only replicate in a single model system. The potency of this targeting can be visualized by comparing a Sendai virus encoding GFP that includes either a scrambled 3’UTR (denoted SeV-GFP) to one targeted by host miRNAs (SeV-GFP-T). Comparing the viral hemagglutinin-neuraminidase (HN) protein to GFP illustrates how potent miRNA targeting can be in the presence of a functional miRNA system. Despite a complete loss of GFP in wild type cells, fluorescence is visible when SeV-GFP-T infects cells lacking either Dicer (Dcr-/-) or Ago2 (Ago2-/-) – the only Argonaute protein capable of cleaving its target when perfect complementarity is achieved between miRNA and its target.
In addition to vaccines and biocontainment, we use miRNA-based targeting to compare the genetic plasticity of viruses to determine how distinct viral families adapt to the same selective pressure. These efforts not only demonstrate that positive strand RNA viruses undergo rapid homologous recombination to divert targeting, but they also revealed a unique aspect of our own biological defense systems. In contrast to positive strand RNA viruses which rapidly excise the targeting cassette, miRNA-based targeting of negative strand RNA viruses completely eliminated any expression of the virus for up to six days post-infection. However, beyond this point, we observe consistent escape of the virus as a result of multiple A to G substitutions across all of the miRNA targets. This viral escape is lost in cells deficient in the double strand RNA deaminase (ADAR1) and those lacking AGO1/3/4 unless they are reconstituted (Figure 1E). These results suggest that ADAR1 may utilize miRNA biology to perform its recently described function of eliminating double stranded RNA production from the cell.